DEVELOPING MARKER-ASSISTED BREEDING FOR QUALITY AND DISEASE RESISTANCE TRAITS IN BRASSICA OILSEEDS.
Daryl Somers, Gerhard Rakow, Philip Raney, Vinod Prabhu1, Ginette Séguin-Swartz, Roger Rimmer, Richard Gugel, Derek Lydiate,
Andrew Sharpe.
Agriculture and Agri-Food Canada-Saskatoon Research Centre, 107 Science Place, Saskatoon, SK, Canada, S7N-0X2. (email – SomersD@em.agr.ca)
1Division of Genetics, Indian Agricultural Research Institute, New Delhi, 110012, India.
ABSTRACT
Improvements in Brassica oilseeds for both canola quality and disease resistance traits are essential to keep canola oil in high demand on international markets. Over the last three years, the Saskatoon Research Centre has invested substantially in canola by developing and implementing marker-assisted breeding in the canola breeding program. High throughput DNA fingerprinting labs have been established that make use of RFLP, AFLP, RAPD and microsatellites markers as well as genetic maps. These technologies will allow us to create complex genotypes from crosses that segregate for multiple, polygenic traits.
Some specific projects include developing marker-assisted breeding for yellow-seed colour, low linolenic acid, high oil content, and resistance to blackleg and white rust. All of the quality traits are under polygenic control, for example, yellow seed colour is controlled by three loci and linolenic acid is controlled by two loci. In addition, there are multiple sources of resistance for blackleg and white rust. The marker-assisted introgression of these important traits into species such as B. rapa, B. juncea and S. alba is also being explored. For each gene/QTL, PCR based allele specific amplicons are being developed as high throughput, marker-assisted breeding tools. These projects will be discussed with reference to marker technology, plant breeding and gene pyramiding.
KEYWORD: Leptosphaeria, Albugo, fatty acids, seed colour
INTRODUCTION
The Saskatoon Research Centre of Agriculture and Agri-Food Canada (AAFC-SRC) is the ‘centre of excellence’ for canola research in Canada. Aside from some dramatic improvements and expansion to the research facility in recent years, there has also been the addition of a molecular genetics section of researchers. Among this group, there are four scientists that are actively involved in all aspects of molecular marker development and marker-assisted breeding. Working in concert with the oilseed breeding section which includes scientific programs in breeding, chemistry and pathology, the AAFC-SRC is expected to play a significant role in the accelerated breeding of high quality, disease resistant Brassica oilseeds in Canada.
A mixture of marker technologies are being employed by researchers at AAFC-SRC to improve and accelerate the development of elite germplasm for use by the Canadian plant breeding industry. There is a substantial emphasis on unique quality traits and novel disease resistance that AAFC-SRC expects will help maintain the value of Canadian canola oil and meal on international markets. The high quality traits extend beyond the current canola species, Brassica napus and Brassica rapa, and into potential future canola quality mustard species such as Brassica juncea, Sinapis alba and Brassica carinata. The objective of this report is to describe available marker technologies and to review the development, approaches, and strategies for marker-assisted breeding in Brassica oilseeds at AAFC-SRC. A few examples of traits that are currently under study and where marker-assisted breeding is being used are presented.
TECHNOLOGIES
Restriction fragment length polymorphism (RFLP)
In the past, and currently, genetic maps are developed using RFLP markers which rely on homology of a DNA probe, typically 200 to 1,000 bp in length, to chromosomal loci. Since this technique is homology based, RFLPs have a strong ability to detect multiple, homologous loci. In addition, DNA probes from related genera can be hybridized to a variety of plant genomes and thus, there is a large collection of DNA probes available to construct a genetic map. This is particularly useful in crucifer genetics where DNA probes from Brassica and Arabidopsis can easily be cross hybridized to an alien species. Typically, there is a good degree of synteny between the marker or gene order in related crucifer species.
Aside from constructing genetic maps, RFLP loci have been extremely useful in comparative mapping studies where a common set of probes can be used to make genetic maps in related crucifer species. The common set of homologous RFLP loci then allows researchers to align maps from related species. Our labs are currently interested in creating maps in crucifer genera such as Sinapis, Eruca, Moricandia and Raphanus that carry unique traits. When the loci are mapped using Brassica DNA probes, the marker-assisted introgression of an alien chromosome interval is detectable in hybrid progeny and backcross progeny.
Random amplified polymorphic DNA (RAPD)
RAPDs have often been described as markers which are not reproducible, and less polymorphic than other types of markers. Although RAPD markers are less polymorphic, they can be highly reproducible in Brassica oilseed species. We have made extensive use of RAPDs in the past to perform gene tagging experiments and will continue with this technology in the future. RAPDs can be used to tag simple, dominant genes as well as identify multiple chromosome intervals controlling a quantitative trait. The limitation of RAPDs imposed by the lack of molecular polymorphism can be overcome by developing methods to screen RAPD markers and primers in high numbers. We have identified a set of 350 RAPD primers from the UBC (University of British Columbia) collection that amplify robust and reproducible bands in oilseed Brassica species. There is an average of seven DNA fragments amplified from each primer, which translates into over 2,400 alleles that can be amplified and analysed in a gene tagging experiment. When this capacity is coupled with high throughput PCR-based fingerprinting techniques, it is very efficient both in cost and time to identify RAPD markers associated with a trait. This is exemplified below in both quality and disease resistance projects.
Amplified fragment length polymorphism (AFLP)
The AFLP technique can resolve allelic length polymorphism’s at 20-60 loci simultaneously in Brassica oilseeds and the capacity to detect polymorphism is good. There are three primary applications of the AFLP technology in marker-assisted breeding labs. The first is to generate genetic maps that can be used to perform QTL analysis in segregating populations. The second application is to use AFLP technology with bulked segregant analysis (BSA) to identify DNA markers near genes controlling traits of interest or to saturate an interval of a chromosome with markers to define the position of a gene. The third application is to carry out accelerated backcrossing experiments by creating DNA fingerprints from well spaced loci over the whole genome in backcross progeny.
AAFC-SRC is using two detection methods that include 33P labeled primers and autoradiography as well as fluorescent labeled primers and detection on an ABI 377 system. In both cases, when a sufficient amount of equipment is available, the capacity to perform extensive BSA on a trait or to generate marker segregation data in the development of genetic map is greatly enhanced.
Microsatellites
Microsatellites are short tandem repeats of 2 to 4 nucleotides in length that are highly dispersed in the genomes of plants, animals and microorganisms. Allelic variation at a microsatellite locus is detected by PCR amplifying the segment of DNA containing the microsatellite; different genotypes will have different copy numbers of the repeat and thus length variation. A consortium of 14 plant breeding companies including AAFC-SRC has initiated a project to develop 1,500 microsatellite markers in B. napus and to create a genetic map in an established mapping population. This initiative will develop a high density map of PCR based markers (microsatellites) which can be aligned to existing Brassica genetic maps, similar to those developed at the John Innes Centre, Norwich, UK.
The advantage to this high density map is the capacity to analyse very complex, segregating populations, or to make populations with multiple parents and be able to reassemble/select individual plants with complex yet specific genotypes. The technology is all PCR based and several robotic work stations assist with the large number of DNA samples and PCR reactions to be handled. This high throughput DNA analysis strategy using microsatellites will enhance the capacity to carry out accelerated backcrossing to restore a desirable genetic background now carrying novel traits.
Allele specific amplicons (ASAs)
Candidate genes with known functions can be PCR amplified in segregating populations and subsequently mapped to an interval known to control a trait of interest. The gene can then act as a specific sequence template that is associated with the trait. If PCR primers can be designed, based on the gene sequence, to specifically amplify the allele controlling the trait, then this is referred to as an allele specific amplicon (ASA). It is not always necessary to use a candidate gene as the sequence template to design the ASA primers. It is also possible to use random markers such as a RAPD, AFLP or RFLP marker that is tightly linked to the gene(s) controlling the trait.
Often the sequence polymorphism between mutant and wildtype alleles can be quite subtle, possibly only a single base pair difference. It is possible to exploit these differences in PCR primer design, typically by incorporating the sequence difference in the 3’ end of the primer. Extensive testing of the ASA primers is then required which may include altering the Mg concentration and annealing temperature used in the thermal cycling protocol. We have found that using “touchdown” PCR with particular attention to annealing temperature and number of cycles can have a dramatic, positive effect on the outcome of a developing ASA test.
Population development
The ability to regenerate fertile plants from haploid microspores in certain Brassica species has had a major impact on genetic studies and plant breeding. Doubled haploid (DH) lines are homozygous at all genetic loci and thus allows a breeder to “fix” a genotype in a true breeding state and immortalizes this fixed genotype in all lines by maintaining selfed seed of each line. In genetic and molecular mapping studies, we have used microspore culture to generate segregating populations arising from complex crosses. A large number of traits each controlled by multiple genes can be captured in a DH population and the population is then phenotyped over multiple years and in two or three locations.
The other advantage of DH populations is the capacity to analyse traits within the population using bulked segregant analysis (BSA). This genetic technique requires that bulked DNA samples be prepared from individual lines sharing a common phenotype, typically extreme phenotypes (Michelmore et al. 1991). The DH population technology enables the bulked DNA samples to be derived from only homozygous individuals. Thus each bulk is “pure” for a contrasting allele at a segregating locus that controls a portion of the phenotypic variation and these differences are more easily identified in screening random molecular markers for polymorphisms.
In situations where a DH population is difficult to develop, we have resorted to single seed decent (SSD). Oilseed species such as B. napus and B. juncea respond well to accelerated growth conditions which include small pot size, high light intensity, long day length (18-20 hrs), elevated growth temperatures (24°C) and constant pruning of leaves and secondary racemes. When seedlings are grown under the above conditions, the plants may be forced to flower in just 2-3 weeks and mature in 12 weeks. Thus an F2 population can be carried forward to the F6 (inbred) generation rather quickly and enter a field analysis as F6 rows, which could be considered to be homozygous at almost all loci. The advantage of SSD over DH populations, is having more control over the final population size. We have even attempted SSD with Sinapis alba, an outcrossing species; an initial 500 F1 seedlings was reduced to 150 F4 plants as a result of inbreeding depression on fertility. Although the number of lines has been drastically reduced, the population is still suitable for field evaluation of quantitative traits and we have a good prediction of how to proceed with any future SSD populations developed in S. alba.
QUALITY TRAITS
Low linolenic acid in oilseed Brassica
There is a substantial effort by both private and public breeding programs to develop novel fatty acid traits in Brassica oilseeds, namely a high oleic, low linolenic acid profile. The typical B. napus canola oil profile is 65% C18:1 (oleic acid), 20% C18:2 (linoleic acid) and 10% C18:3 (linolenic acid). The development of low C18:2 and C18:3 canola types indirectly raises the C18:1 levels and produces a canola oil with greater heat stability (high C18:1) and reduced potential to go rancid (low C18:3). This new novel fatty acid profile of B. napus would be >80% C18:1, <10% C18:2, <2% C18:3. Breeding for low C18:3 levels is challenging since the genes are inherited in a recessive manner, often the novel phenotype is a result of mutagenesis (Röbbelen and Nitsch 1975, Rakow 1973) as in the varieties Apollo and Stellar (Scarth et al. 1988, Scarth et al. 1994). Also, the low C18:3 trait is influenced significantly by the environment, which makes selection of plants by bulk seed or half seed analysis in the greenhouse unreliable.
Several papers are published where markers are identified that are associated with QTLs that control the phenotypic variation in C18:3. In summary, it appears that there are two major loci controlling the trait and one of these encodes a FAD3 gene (Somers et al. 1998, Jourdren et al. 1996). This FAD3 gene is responsible for desaturation of C18:2 to C18:3 in the microsomal cell fraction. The challenge now is to develop ASA type markers for both of the important loci and to implement marker-assisted selection for Brassica oilseeds such as B. napus, B. rapa and B. juncea. We are currently experimenting with marker-assisted selection of the two B. napus loci controlling low C18:3 in a canola quality B. juncea background. There are two streams of selection where plants are 1) selected in a conventional way through chemical analysis or 2) selected only on the basis of the DNA fingerprint. The molecular strategy uses successive, accelerated backcrossing making selections based on gene tags as well as AFLP analysis of the whole genome. For example, in 129 BC2F1 seedlings, there were 31 seedlings selected that carried both B. napus loci for low C18:3. Among these 31 BC2F1 seedlings, there was considerable variation in the amount of B. napus genome present. The conventional method includes keeping the populations large and selfing selected half seeds in the greenhouse to check the bulk seed sample for low levels of C18:3. There is a marginal correlation between which plants are selected by chemical analysis of half seeds and which are selected by the low C18:3 gene tags. The molecular approach will result in BC4F3 rows to be tested in the field in year 2000 where the success and efficiency of this selection strategy will be evaluated.
An AFLP map is being generated in a DH population that segregates for linolenic acid content as well as seed colour, a few of the developing linkage groups are presented in figure 1. The FAD3 gene is shown on linkage group A and a second, presumably FAD gene, is located on linkage group B. The high density of markers in these two regions is the result of targeting markers to the intervals by BSA. Markers with the prefix “LA” or “YN” are RAPD markers, all others are AFLP markers. In this population, these 2 loci explain 48% of the phenotypic variation in C18:3 levels.
Yellow seed colour in B. napus
The same DH population described above, derived from Apollo X YN90-1016, segregates for seed colour. In fact the segregation is transgressive with the parents showing whiteness index readings of 1.0 and –13.0 respectively, whereas there are DH lines that have whiteness index readings of –23. The DH lines were field tested for 2 years and the seed colour frequency distribution was clearly bimodal. Two attempts at seed colour gene tagging with BSA were performed, the first strategy used DNA bulks made from 10 black-seed lines and the 10 best yellow-seed lines. The second strategy used the 10 best yellow-seed lines and the 10 worst yellow-seed lines. In the first strategy, several RAPD markers were identified that tagged a qualitative, Mendelian gene that controlled whether a plant would produce black seed vs yellow seed. The gene was the genetic element responsible for the bimodal distribution and was tentatively called Pigment 1 (Fig 1., linkage group D). The second strategy identified two additional QTLs that controlled the degree of yellowness in lines grouped in the yellow seed mode of the distribution. Together, the two yellow-seed QTLs explained approximately 27% of the variation in yellow seed colour. The QTLs are positioned in figure 1 on linkage groups B and C centered on markers YN12 and YN9 respectively. Note the close proximity of YN12 to the unidentified FAD gene, the favourable alleles controlling yellow seed colour and low linolenic acid are tightly linked in repulsion in this cross. A related genetic study has shown that yellow seed colour is expressed when all three genes/QTLs are in a homozygous recessive state. All of this data is consistent with a study by Shirzadegan (1986) who developed a model of three recessive genes, where gene Bl1 was epistatic over two QTLs and controlled the presence and absence of seed pigment.
At AAFC-SRC, Dr. Gerhard Rakow has made significant advances in the development of yellow-seeded B. napus where the seed colour is more environmentally stable, the yellow-seeded lines have decreased fibre content and greater oil content over black-seeded types, in isogenic comparisons (Rashid and Rakow, 1995). The yellow-seeded B. napus requires some substantial improvements in disease resistance and improvements towards superior, novel fatty acid profiles. The markers we have developed for the yellow-seed trait in B. napus will play an important role in re-selecting the good yellow seed colour in segregating populations.
Figure 1. A partial genetic map based on AFLP and RAPD markers that shows linkage groups that carry genes/QTLs that control seed colour and linolenic acid content in B. napus. Markers with the prefix “LA” or YN” are RAPDs and all others are AFLP markers.
 |
Oil content in B. napus
Another important quality improvement in Brassica oilseeds studied in the lab is selection to increase oil content in B. napus, and B. juncea. We are focusing on a DH population segregating with a >10% range in oil content. The DH lines, on a row basis, contain from 35% to 48% oil. This population has been phenotyped with over five location years at the end of 1999 and is being analysed by BSA to identify markers associated with the extremes in oil content. The extent to which the population is field tested exemplifies the difficulty with selecting high oil lines from segregating populations. Both locations and years can have a dramatic effect on oil content, and this trait cannot be evaluated reliably in the greenhouse. Therefore, the development of marker-assisted selection tools for QTLs controlling oil content is important in order to provide breeders with the opportunity to simultaneously focus on other novel quality and disease resistant traits all segregating in the same, large complex population.
Currently, we are developing two populations with >300 lines each, of B. juncea that segregate for oil content with a range >10%. One population segregates for erucic acid content and the other population is in a zero erucic acid background. From these populations, we can observe and interpret the effects of erucic acid on QTL analysis for oil content in B. juncea. These populations are going through SSD and will be ready for evaluation at two locations in the year 2000 at the F6 generation.
DISEASE RESISTANCE
Blackleg resistance (Leptosphaeria maculans)
An emerging set of data in our labs suggest that there are at least two genes located in a small interval of the B. napus genome that provide resistance to western Canadian isolates of L. maculans. Preliminary data suggests there may be a distinct adult plant resistance and cotyledon stage resistance gene. We are using several DH populations where Westar is the susceptible parent and the resistant parents include Shiralee, Maluka, Cresor and RB87-62. These blackleg resistant parents show different types of resistance ie. Cresor has good adult stage resistance and little if any cotyledon stage resistance, the Australian lines appear to carry both types of resistance and recombinants can be found between these phenotypes. We have developed two closely linked markers (3 cM apart) that flank the adult stage resistance gene in Cresor. These RAPD markers have been converted to ASA type markers and appear to be polymorphic between most B. napus lines that differ in resistance/susceptibility to blackleg. The markers are likely to be useful in selecting for cotyledon stage resistance as well because of the close proximity of this gene to the markers. There is some suggestion from our phenotypic and molecular data that there is little genetic variation for blackleg resistance at this locus and that alternate sources of resistance mapping to different locations should be explored to develop more durable blackleg resistance in B. napus and B. rapa.
White rust resistance (Albugo candida)
Recent research in our lab has shown there may be two genes that provide resistance to Albugo candida (race 2A) in the AAFC-SRC B. juncea breeding program. One of the genes, Ac2a1 is dominant and molecular mapping of this gene was described by Prabhu et al. (1998) and Cheung et al. (1998). By surveying a collection of B. juncea with both similar and related pedigrees, the source of the resistance and PCR based markers was traced to the Russian line Donskaja, which was introduced to the AAFC-SRC breeding program as a high oil content B. juncea line from the Vavilov Breeding Institute in Russia. We have made significant advances in developing ASA type markers from RAPD bands shown to map in flanking positions to the dominant resistance gene Ac2a1 (Prabhu et al. 1998).
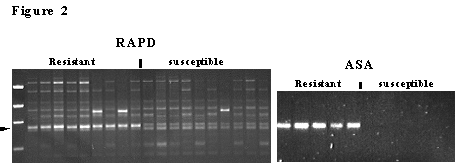
One example of an ASA test for a marker closely linked to Ac2a1 is shown in figure 2. The marker was originally identified in a DH population using BSA and was mapped 1.4 cM from the Ac2a1 gene. The RAPD marker was cloned, sequenced, and primers were designed to specifically amplify the allele in resistant lines only. This ASA marker and others are polymorphic in several B. juncea lines, particularly the lines used in the canola quality B. juncea breeding program, where this resistance and associated markers will be most useful.
Figure 2. Agarose gels showing the PCR amplification of a RAPD marker linked to the B. juncea white rust resistance gene Ac2a1, that was converted to an ASA. The RAPD marker maps 1.4 cM from Ac2a1 and the ASA was develped by cloning the RAPD marker and designing a specific pair of primers for this amplicon. The markers are only amplified in white rust resistant lines.
A second gene, Ac2a2 was identified in a cross of ACVulcan x Common Brown. This resistance appeared to be recessive or partially dominant, where the F1 showed an intermediate level of resistance, between the parents. The cross used to tag the Ac2a2 gene appears to be cytologically unstable and new crosses have been developed to accurately map and tag this resistance gene.
Finally, the canola quality B. juncea breeding program has used an interspecific cross of B. juncea X B. napus to introgress certain B. napus fatty acid profiles and glucosinolate content. The backcross progeny in this program have shown a good resistance to A. candida (race 2V), clearly arising from the B. napus parent. Several populations are being considered for tagging this introgressed segment in B. juncea.
The three resistance genes should prove to be useful in both condiment mustard breeding and canola quality mustard breeding. We intend to pyramid the resistance genes together using markers in B. juncea to provide a robust, horizontal type of resistance to A. candida.
SUMMARY
There have been significant advances in molecular marker technology beginning with RFLP, Southern-based markers, to PCR-based markers such as microsatellite markers. A driving force has been the need to develop technology that permits high throughput marker analysis toward rapid trait introgression and accelerated backcrossing. Plant breeding has become far more competitive and molecular marker tools are enhancing the ability of plant breeders to create unique genotypes, faster. Molecular marker technology does not replace the skill of visual observation and chemical analysis that plant breeders make use of today; it is a tool that should enhance or augment current plant breeding practices. Important, complex traits in canola such as yield, hybrid vigor, stress tolerance and adaptation are not selected for with markers, but plant breeders may be able to focus more attention on these complex, quantitative traits and use markers to select for more simply inherited, novel traits or use markers to select for a recurrent background.
There is an emphasis in the AAFC-SRC oilseed breeding and molecular genetics programs to identify new genetic variants for quality characteristics and disease resistance. If these new genetic variants can be characterized by molecular mapping and gene tagging, then their introgression into elite, high yielding canola germplasm can likely be accelerated. Molecular breeding has moved almost entirely toward PCR-based technology such as AFLP, ASA and microsatellite markers. These technologies are amenable to automation and thus accuracy and high throughput.
ACKNOWLEDGEMENTS
We wish to thank the following people at AAFC-SRC for assistance with all the research described in this paper; Goewin Demmon, Jason Danielson, Jennifer Helston, Ken Friesen, Hossein Borhan, Bin Zhu and Todd Olsen. Financial support was provided by Saskatchewan Agriculture and Food-Agriculture Development Fund, Canada-Saskatchewan Agri-Food Innovation Fund.
REFERENCES
Cheung, W.Y et al. (1998) Identification of RFLP markers linked to the white rust resistance gene (Acr) in mustard (Brassica juncea (L.) Czern. and Coss.). Genome 41:626-628.
Jourden, C. et al. (1996a) Specific molecular marker of the genes controlling linolenic acid content in rapeseed. Theor Appl Genet 93:512-518.
Michelmore, R.W. et al. (1991) Identification of markers linked to disease resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc. Natl. Acad. Sci. 88:9828-9832.
Prabhu, K.V. et al. (1998) Molecular markers linked to white rust resistance in mustard Brassica juncea. Theor. Appl. Genet. 97:865-870.
Röbbelen, G. and Nitsch A. (1975) Genetical and physiological investigations on mutants for polyenoic fatty acids in rapeseed, Brassica napus L. Z PflanzenzÜchtg 75:93-105.
Rakow, G. (1973) Selektion auf Linol- und Linolensauregehalt in Rapssamen nach mutagener Behandlung. Z PflanzenzÜchtg 69:62-82.
Rashid, A. and Rakow G. (1995) Seed quality improvements in yellow seeded Brassica napus. Ninth International Rapeseed Congress, July/1995, Cambridge, UK, pp 1144-1146.
Scarth, R. et al (1988) ‘Stellar’ low linolenic-high linoleic acid summer rape. Can J Plant Sci 68:509-511.
Scarth, R. et al. (1994) Apollo low linolenic acid summer rape. Can J Plant Sci 75:203-204.
Shirzadegan, M. (1986) Inheritance of seed colour in Brassica napus L. Z. Pflanzenzuchtg 96:140-146.
Somers, D.J. et al. (1998) Identification of molecular markers associated with linoleic acid desaturation in Brassica napus. Theor. Appl. Genet. 96:897-903.